library(Hmisc)
# First read in the temperature data, and assign column names.
temperatureDF <- read.table("data/temp.dat", col.names = c("Temp", "Time"))
temperature <- temperatureDF$Temp
time <- temperatureDF$Time
# We now want to regress our data against sinusoids with a period of 72.
# First create the sinusoids.
cosine <- cos(2 * pi * time/72)
sine <- sin(2 * pi * time/72)
# Then fit a linear model.
lm.temperature <- lm(temperature ~ cosine + sine)
temperatureResiduals <- lm.temperature$residuals
x<-matrix(0, nrow = 350, ncol = 2)
y<-temperatureResiduals[3:352]
for(i in 1:2){
x[,i]<-Lag(temperatureResiduals, i)[3:352]
}
ar.reg<-lm(y~x-1)
summary(ar.reg)
par(oma = rep(2, 4))
par(mar = c(5, 5, 1, 1))
par(mfrow=c(2,1))
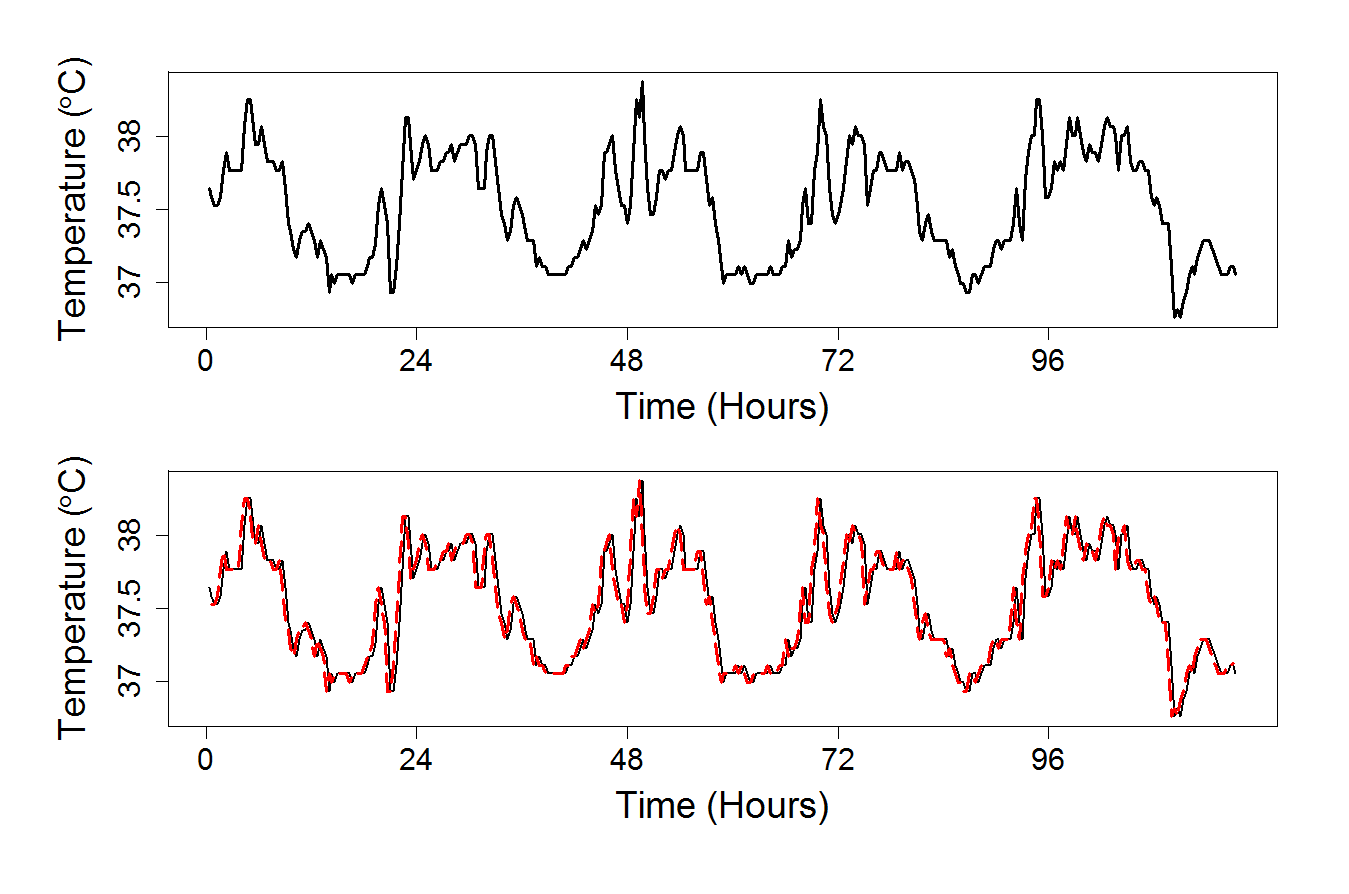
plot(temperature, type="l", xaxt = "n", yaxt = "n", xlab = "Time (Hours)", ylab = expression(paste("Temperature (",degree,"C)")), main = "", lwd = 3, cex.lab = 1.6, col = "darkgrey")
axis(1, at = seq(from = 0, to = 288, by = 72), labels = seq(from = 0, to = 96, by = 24), cex.axis = 1.5)
axis(2, at = seq(from = 36.5, to = 38.5, by = 0.5), labels = seq(from = 36.5, to = 38.5, by = 0.5), cex.axis = 1.5)
plot(temperature, type="l", xaxt = "n", yaxt = "n", xlab = "Time (Hours)", ylab = expression(paste("Temperature (",degree,"C)")), main = "", lwd = 2, cex.lab = 1.6, col = "darkgrey")
lines(2:351, lm.temperature$fitted[3:352]+ar.reg$fitted, col = 2, lwd = 3, lty = 5)
axis(1, at = seq(from = 0, to = 288, by = 72), labels = seq(from = 0, to = 96, by = 24), cex.axis = 1.5)
axis(2, at = seq(from = 36.5, to = 38.5, by = 0.5), labels = seq(from = 36.5, to = 38.5, by = 0.5), cex.axis = 1.5)
dev.print(device = postscript, "18.8.eps", horizontal = TRUE)