Covid-19 Science Resources
Website created April 13, 2020; last updated April 22, 2020.Here are some of my personal recommendations for The Latest Science, Popular Press, Specialized Sources, Published Papers, following a few comments.
Important updates to this webpage (April 19): (i) revision of my comments, with a link to an opinion piece criticizing IHME methodology; (ii) a comment on herd immunity; (iii) additional comments, which are more technical, may be found at the end. Also, I am adding links as I become aware of them, but I am trying to keep the whole as brief as I can. As of April 19, NIH's curated publication database had about 1500 papers (link below).
Comments
April 15:
- I am not an expert on any of this! I created the page out of frustration that I could not find any such brief set of suggested resources. I'm not sure how well I'll do at updating it. It would be GREAT if someone else did a better job, and updated it! I would, however, very much appreciate comments and suggestions.
- I am trying to stick mainly to science and avoid policy. Of course, we care about the science in large part for its value in making policy, and there are places where it is interesting to see how the scientific knowledge will be used. An example is the April 17 guide for governors by an expert committee at Johns Hopkins. There is a lot that could be done to better understand and improve the human process of making decisions in the presence of uncertainty.
- If you look at the modeling results in the published papers, you will be struck by the very large uncertainty throughout. It is crucial to keep in mind the large uncertainties in the projections---they are not necessarily "wrong" in what they predict if their estimates are off, even by a fair amount, unless the estimates are off by so much as to deviate beyond what their estimated uncertainty would suggest to be fairly likely. I should add that the reported uncertainties are themselves based on the models and, thus, on the modeling assumptions. As always, even the most informative scientific conclusions should be taken as guides to understanding: a little illumination keeps us from stumbling in the dark, but it can not transport us to our destination. Concerning COVID projection uncertainty see the FiveThirtyEight survey results (below), and the NY Times article on the uncertainties (below).
- Published papers from reputable journals vary in quality, and strength of evidence. Furthermore, a lot of the papers (including some in the links I am sharing) have either not yet been reviewed, or were given expedited review---which suggests it is appropriate to have an extra bit of scientific skepticism about aspects we can't judge for ourselves.
- There are many, many scientific issues surrounding COVID-19 and some of the ones I omit from the web page have interested me enough to take a look. There are, of course, a lot of publications based on lab experiments (which I tend to like), though they are sometimes coupled with observational data. I will mention the number, and relative number, of ACE2 receptors in the lung and heart, in relation to ACE inhibitor blood pressure meds, which I happen to take. Turns out the evidence about ACE inhibitors is mixed (I'm inclined to believe the "protective" results rather than the "increased risk" results). A nice blog entry about the issue is here and a review is here.
- Of particular interest is a key variable (which my physician-wife Loreta and I have talked about a lot): inoculum size. This almost never gets discussed, mainly, I presume, because it's nearly impossible to measure or estimate, except in a laboratory setting. There's a helpful lab study, on humans exposed to H1N1, here. For me, the main findings are (i) inoculum size matters to both probability of getting the disease and severity of symptoms and (ii) there is very large variation across subjects (people). See this figure. Also, there's nice New Yorker article that discusses this and related issues.
- Concerning the projection methods themselves, there are two general approaches. One we might call purely predictive. A very popular website with predictions (projections) is produced by a health metrics organization in Seattle, affiliated at least loosely with the University of Washington, called IHME. Their projections are used by the White House Task Force. The methodology is relatively simple, and may be described as curve fitting. In the original version they assumed the death rate over time in each U.S. state would evolve similarly to the way it evolved in Wuhan. A link to their methodology is given below. Subsequent projections have used more recent data, from places other than China, to form baseline accumulated death curves. This change in baseline, however, can lead to noticeable changes in predictions. Furthermore, as I said, the method is simple (a bit more description is in the additional comments, below), and would seem perhaps too simple for many purposes. For example, the assumed similarity among death rate curves may have trouble accommodating the inhomogeneity across locations, especially as one starts to extrapolate more than a short period into the future. A list of cautions about the IHME model can be found here. The second approach applies some version of a well-established epidemiological framework called SEIR modeling (often in a slightly simpler form called SIR). The acronym comes from the stages (we usually call them compartments) each patient must travel through: Susceptible, Exposed, Infected, Removed (i.e., removed from the susceptible population because they are either recovered or deceased). Here, in addition to deaths, one must consider infection rates, and the model can be complicated. Modern forms are statistical, meaning that particular features of the data are assumed to follow probability distributions; this also allows calculation of uncertainty. See additional comments, below. Predictions by the group at Imperial College have received a lot of attention and are favored by the British government; a link to their paper on predictions for many European countries, and a few non-European ones, is given below. They explicitly assume particular forms for several probability distributions based on data from China (and other experience in the literature), and they consider a more involved set of interventions, modeled as affecting the rate of transmission. Interestingly, IHME projections turned out to produce very much higher predicted deaths for the UK than projections using SEIR-style models. My own inclinations lean toward the more detailed modeling approach, and, from what I gather, I tend to think the chief groups doing the SEIR-style European and UK projections are likely to be somewhat more careful and trustworthy. Also, the first approach can be done differently, and made very much more sophisticated, as implemented by the Delphi group here at Carnegie Mellon. They have done extremely well in the past, winning many national competitions in flu forecasting. For Covid-19 they are relying on different kinds of inputs: human judgment from large numbers of humans (filling out short surveys online); human behavior, such as searches in Google (e.g., for products relevant to cold symptoms or for information about the disease); and some anonymized patient and patient-testing data (via certain providers and labs). As I understand things, they hope to improve immediate prevalence estimation and short-term predictions. Because the group is embedded in the wider disease forecasting community (including through the CDC, which requested their effort on this), their results could feed into others aimed at informing policy.
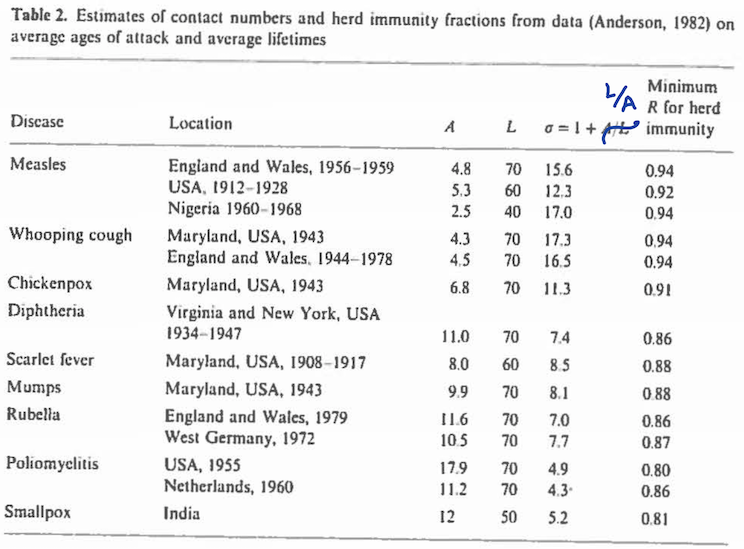
- An extremely important topic is what's usually called "herd immunity." This refers to a stable situation in which every new outbreak will die out, rather than exploding to something unmanageable. From an epidemiological perspective, the main goal is to figure out what percentage of the population needs to become immune, through either exposure or vaccination, in order to reach herd immunity. Here, modeling has been very helpful. I give some links in additional comments, but let me say two things now. First, the simplified theory (using the basic SIR differential equations) provides easily-computed formulas (links below). An informative table for multiple childhood diseases, based on work from the late 1970s and early 1980s (and taken from a 1989 overview found below) may be found here. In that table the lowest percentage immune needed for herd immunity occurs for polio, at 80%. That is, polio becomes manageable once at least 80% of the susceptible population is immune. Other, more highly communicable diseases require higher percentages, e.g., measles requires around 93%. These are, of course, data-based estimates and are subject to uncertainty. The lesson for Covid-19, however, seems clear: because Covid-19 is highly contagious, it would seem at first glance that very high levels of immunity may be required. On the other hand, a second point is that the population of susceptibles is inhomogeneous. The simplest case is if there are two distinct sub-populations, which, in the case of Covid-19, we might imagine to be, say, young and non-compliant (high spreading of the disease) and older, more compliant (lower spreading of the disease). In this case, assuming the young and old interact, but there's much less spreading from young to old than from young to young, the fastest spreading would occur within the young sub-population, and the most important thing to know would be the degree of immunity among the young. This becomes analogous to many childhood diseases. It should provide a more optimistic outlook insofar as immunity among the young could be expected to be much higher than among the old. The literature is full of discussion about situations involving inhomogeneous populations (e.g., see the papers below), but what I've just said is my own speculation. I don't know of good data and modeling to help with analysis of Covid-19 based sub-populations (which would also have to consider special outbreak situations, as in nursing homes), but I presume such data and analysis will come.
- It is important, I think, to realize that the immediate goal of public health efforts is not eradication of the disease but, rather, control of it, where an explosive, widely-spread epidemic is avoided. In my additional comments, below, I mention that the proportion immune needed for herd immunity is based on the assumption that with herd immunity we move to an endemic stage, meaning that the epidemic is gone, but the disease remains in the background. Furthermore, apparently, for the diseases in the cited table immunity is permanent, whereas for the SARS coronaviruses it is likely to taper off across time. This suggests that the proportion immune will itself be dynamic. Also, another unknown is the rate at which this virus might mutate in disease-relevant ways. Endemic Covid-19 would become tolerable if there were treatments that often help patients substantially.
- There many issues associated with vaccine development, as discussed here on March 30 and here on April 9. A technical overview of modern methods may be found here. There are statistical issues, too, and a nice-looking 2009 book by Halloran, Longini, and Struchiner, but I haven't yet found a recent overview.
April 19
April 20
April 22
The Latest Science
National Academy of Medicine >Nature Medicine Click on ``Covid-19 Research in Brief" Note: other specific journals are also good, e.g., JAMA, Science, Lancet, and don't forget the CDC webpage! Also, NIH has a curated, searchable database of Covid papers.
Popular Press
fivethirtyeight.com shines in this context. They are reporting on a survey of expert opinion (a survey of knowledgeable epidemiologists conducted by someone at U Mass), which is helpful due to the very large uncertainties. The April 2 article is here.
See also Nate Silver's April 4 article.
Vox is also doing a nice job. See the pair of April 10 articles on difficulties of prediction and on plans for reopening. Related to reopening: on April 17, an expert committee at Johns Hopkins released a guide for governors. Also on April 17 Science magazine published a short overview news article on next steps.
A nice April 7 article in the NY Times features a professional friend, David Spiegelhalter, who is especially good about public communication, and also Neil Ferguson, head of the Imperial College group. More may be found in David's April 12 overview article from The Guardian, written with Sylvia Richardson.
An April 13 overview on immunity, in the NY Times, by a leading epidemiology modeler (whom I don't know, but who is a colleague and collaborator of an old friend) is here.
A nice graphic showing why Covid-19 is so frightening may be found here and another (from NY Times, April 21) is here, though the latter is part of a "missing deaths" story.
Specialized Sources
Very nice ongoing data summaries.
The IHME model, which is popular and is used by the White House Task Force, but involves very simple curve fitting, is described and here. In contrast, a sophisticated SEIR modeling approach is taken by the Imperial College group.
I provide links to some modeling resources, including tutorials, below (additional comments).
Carnegie Mellon's excellent flu forecasting group (led by two of my close colleagues) has come online with COVID now-casting, with forecasting coming.
Here is a discussion of available R code related to Covid.
As always, a lot of interesting statistical discussion at Andrew Gelman's blog.
Published Papers
Just a few, done by excellent researchers. An early (influential) one here.
Another by the same group, April 8.
And one by the Imperial College group, March 30.
An interesting simulation study that's relevant to policiesfor living with Covid (or "re-opening") is reported and discussed in this article in Science, published April 14.
Additional Comments
More on projections. In the the original version of IHME they (i) fit a curve to summarize the trend in accumulated deaths, across time, in Wuhan, and then, (ii) focusing on the time from social distancing until the curve's rate of increase (the death rate) starts to decrease, assumed the analogous curve for any particular U.S. state would be similar (after adjusting for the age distribution in the local population), and fit such curves to all of the states that had sufficient data (which was most of them, but not all). They used what we statisticians call a hierarchical model, which allowed them to compute measures of uncertainty.
An interesting statistical aspect of the Imperial College work of March 30 is that, rather than estimating all parameters (unobserved variables) from current data, they fix some of them by a combination of judgment and past data. Personally, given the situation, I rather like that idea, as long as it can be justified reasonably well, and as long as it is clear how sensitive results are to the choices.
For those who might care, while the IHME and Imperial College methods are very different, they are both Bayesian.
More on SEIR modeling. For an overview of SEIR modeling, I like this 2018 summary and also a pair of older papers by Hethcote, who himself contributed to the field, one from 1989 and one from 2000. Another nice paper is this 2017 historical overview. There are two related aspects of the work I would call to your attention. First, the models are implicitly probabilistic but too often (for my statistician's taste) not explicitly so. The first appendix in the 2018 summary at least starts down this path. Full-fledged versions are often labeled "stochastic" as in this 2017 primer. Second, when we move from analytical and simulation models to data-based estimation, we need statistical models, in which the data are assumed to follow probability distributions, and this allows investigators to generate assessments of uncertainty. The January 31 paper and the March 30 paper mentioned above are examples. The journal Statistical Science in 2018 had a special issue devoted to inference for infectious disease dynamics (v. 33, no. 1). See the Introduction to that issue.
More on herd immunity. Concerning herd immunity, I would recommend the Hethcote papers from 1989
and from 2000. The starting point is the
notion that herd immunity generates a steady state, or endemic, disease where it can be present at background levels but does not become
an epidemic. The derivation of the key expression for contact number, which leads to the percentage needed for herd immunity, is based on finding
a stable solution to the differential equations; it appeared in a 1975 paper by Dietz,
see Equation (18). Here is a key table from the Hethcote 1989 review, based on data that had appeared in papers by Anderson and May a few years earlier.